Absolute and Simultaneous Quantification of Specific Pathogenic Strains and Total Bacterial Cells
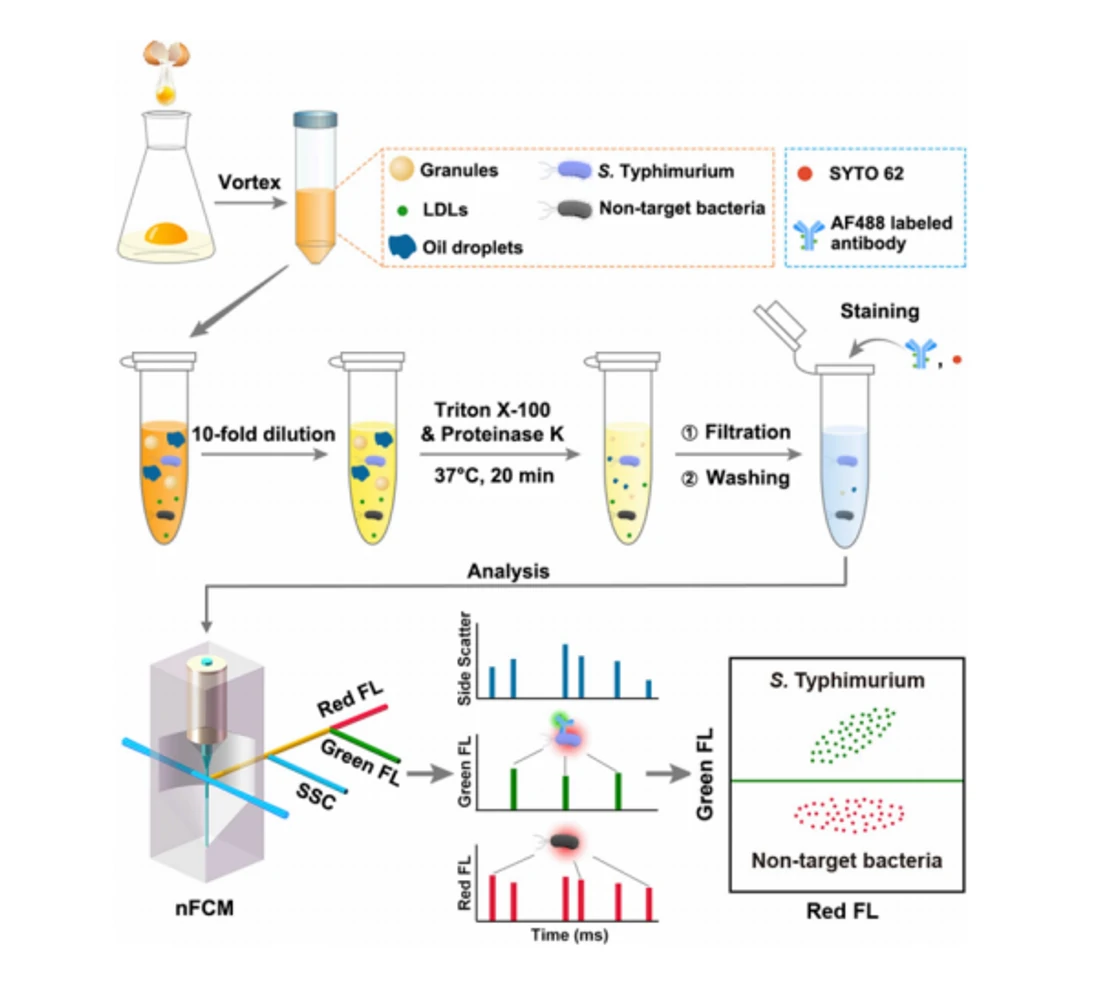
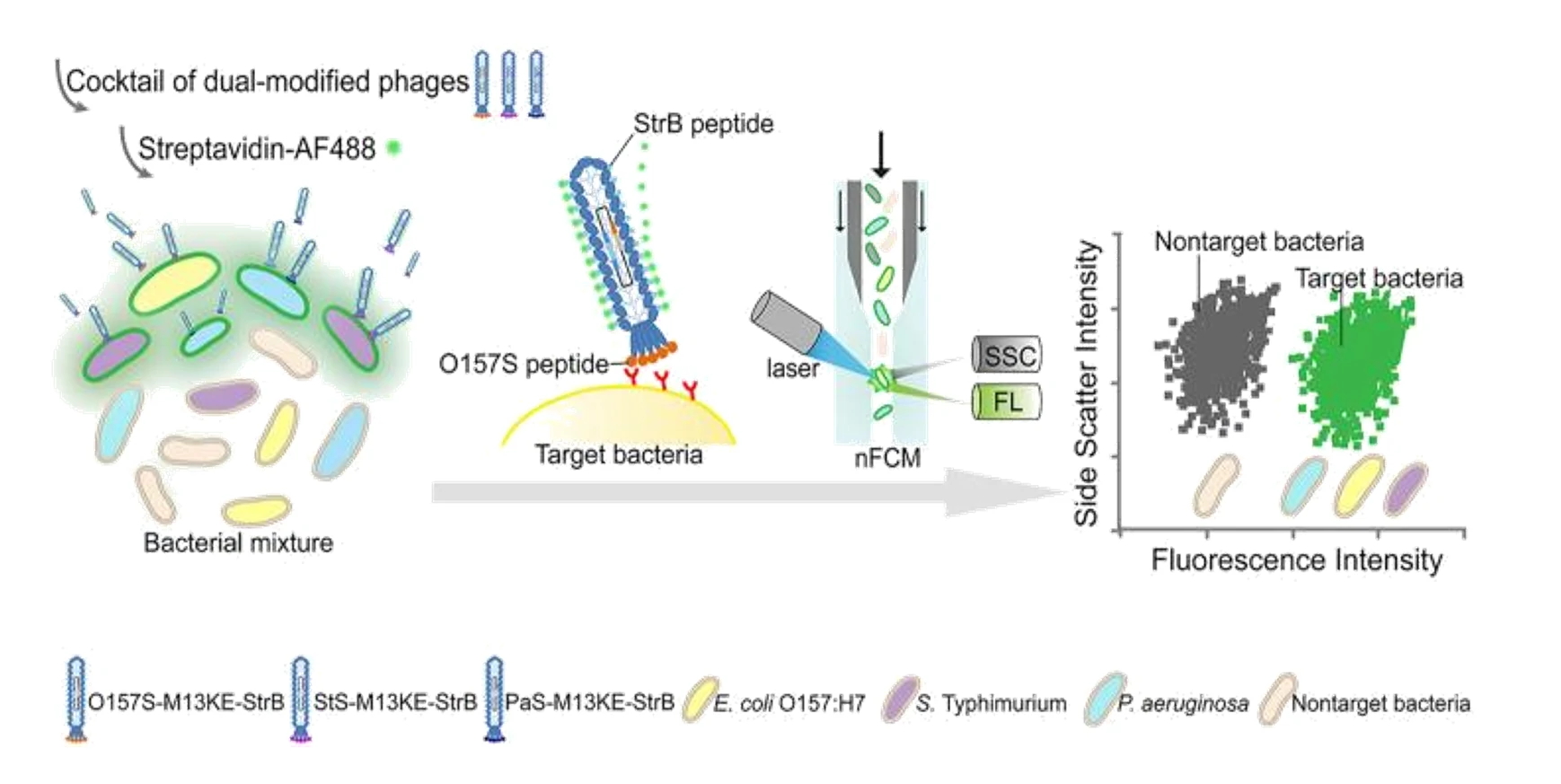
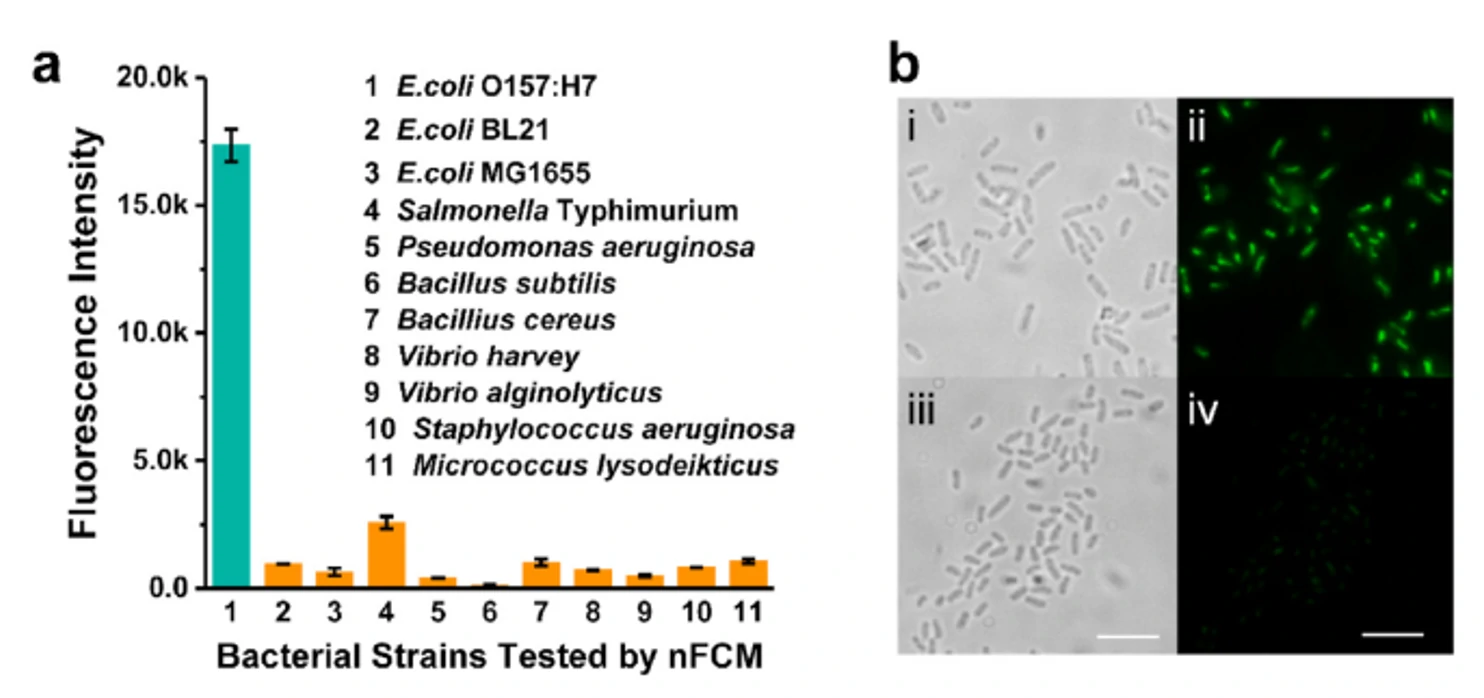
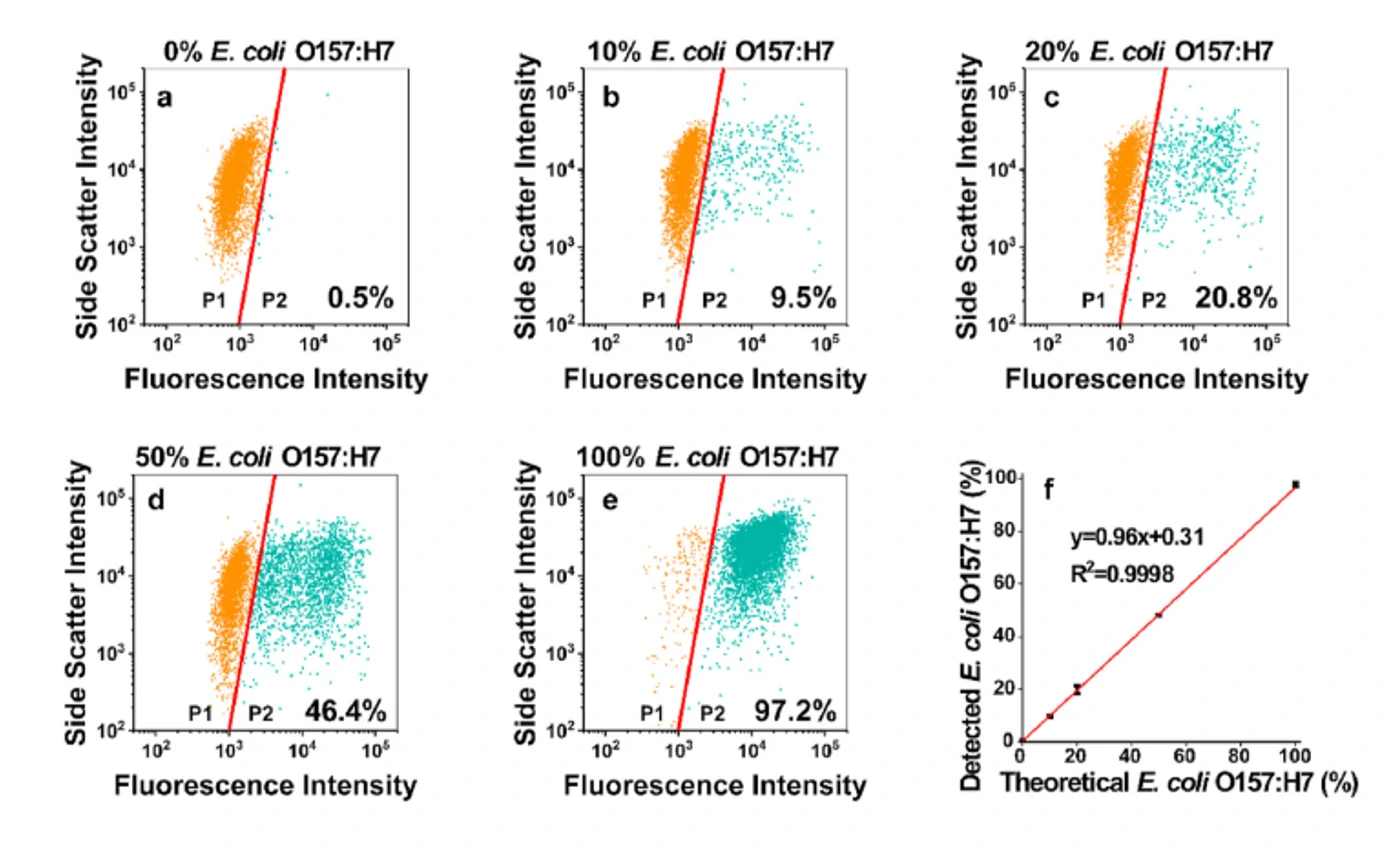
Fast, sensitive, and specific detection of pathogenic bacteria is of great significance for food safety, environmental monitoring, disease diagnosis, treatment, and prevention of bioterrorism. Traditional pathogenic microbial detection methods require the isolation, culture, and a series of biochemical reactions of bacteria, which are complex in operation, long in detection cycle, and cannot be applied to pathogenic bacteria that are difficult to culture. This is far from meeting the actual needs. Traditional flow cytometry still has quite limitations in the highly sensitive detection of pathogenic bacteria. Using Alexa Fluor 647-R-PE as a fluorescent probe for E. coli O157:H7 monoclonal antibody and fluorescent SYTO 9 dye to stain the nucleic acids of bacteria to achieve simultaneous detection of dual fluorescent channels using nanoflow cytometry. Pathogenic bacteria can produce fluorescence signals simultaneously in the green and red fluorescence detection channels, while non-pathogenic bacteria only produce signals in the green channel.
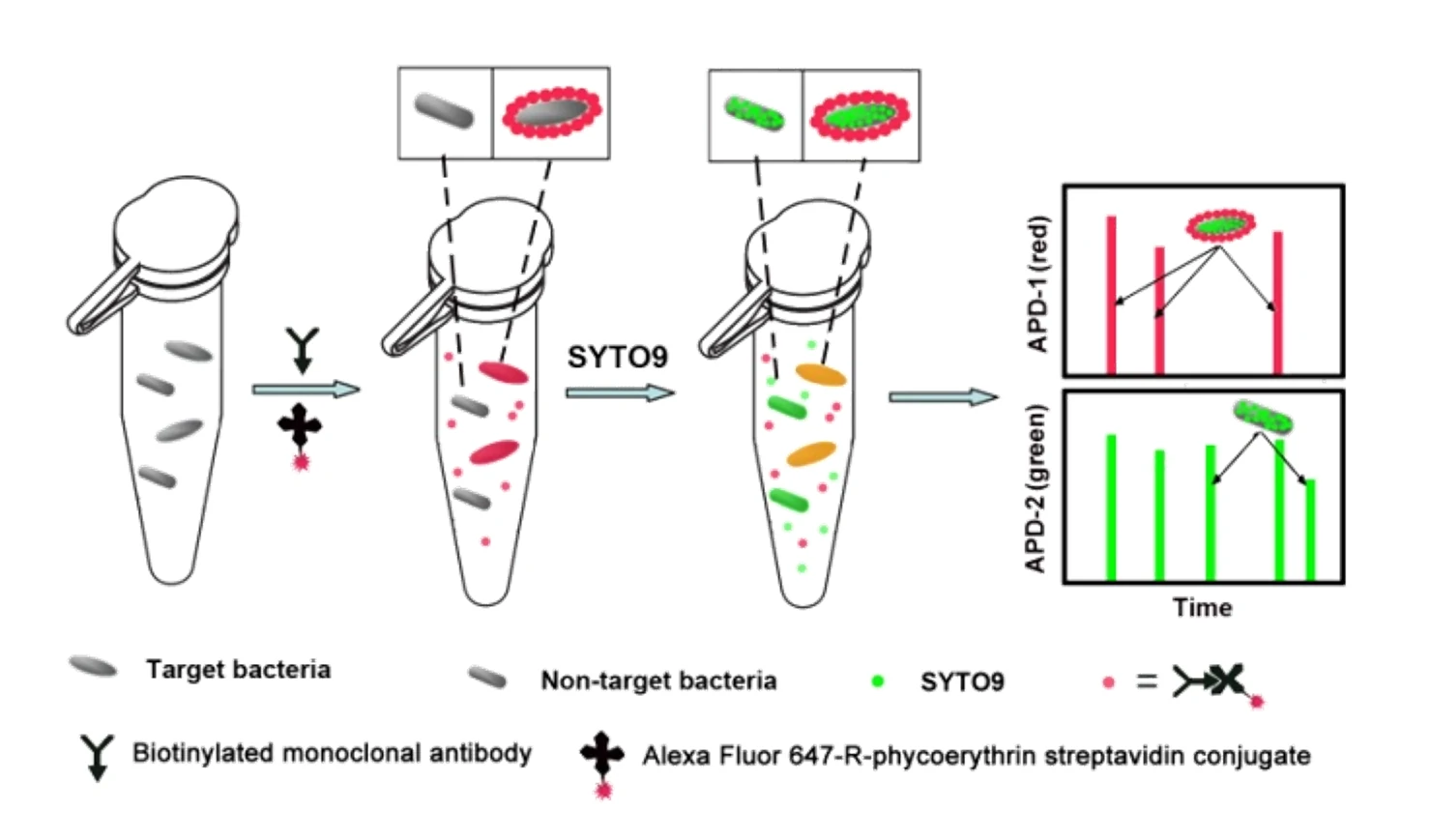
The Flow NanoAnalyzer overcomes the limitations of traditional detection methods, showing potential in the rapid identification and quantitative analysis of pathogenic bacteria in food, environment and disease-related samples.
Anal. Chem., 2010, 82(3), 1109-1116.