Phage cocktail is used for multi-detection of bacterial pathogens
Author: admin Date: February 22, 2024
The Public health sector has an urgent need for rapid, quantitative, and sensitive detection methods for the multi-detection of bacterial pathogens. In this study, the target peptide-specific recognition of pathogenic bacteria was demonstrated through the double modification of the M13KE phage pair. The target peptide was modified on the secondary capsid protein pIII, and the streptavidin-binding (StrB) was modified on the major capsid protein pVIII. The peptide-targeted peptide could specifically recognize the target bacteria, and after binding to the fluorescently labeled streptavidin, the StrB peptide could effectively amplify the signal. With the Flow NanoAnalyzer and fluorescence microscopy, bright fluorescence from individual pathogens can be clearly distinguished from the background.
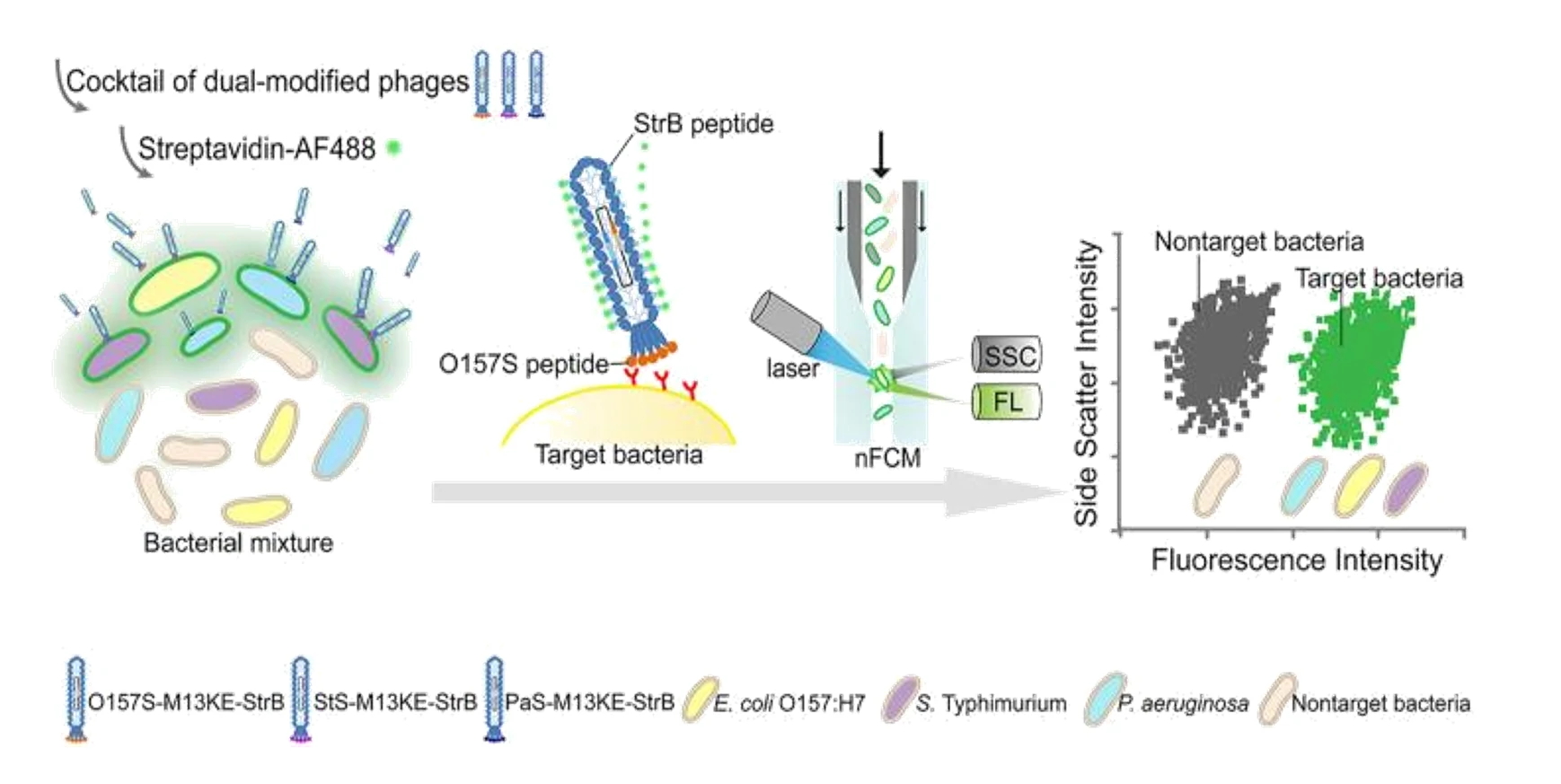
Bacteriophages targeting Escherichia coli O157:H7, Salmonella typhimurium, and Pseudomonas aeruginosa were constructed, and high specificity was verified by a large excess of other non-target bacteria. By analyzing using the Flow NanoAnalyzer, the detection limit of the target pathogen was about 10^2 cells/mL in the presence of other non-pathogenic bacteria with only a 40 mL sample. The combination of three dual-modified phages into a cocktail can also quantitatively and linearly detect three target bacterial pathogens simultaneously.
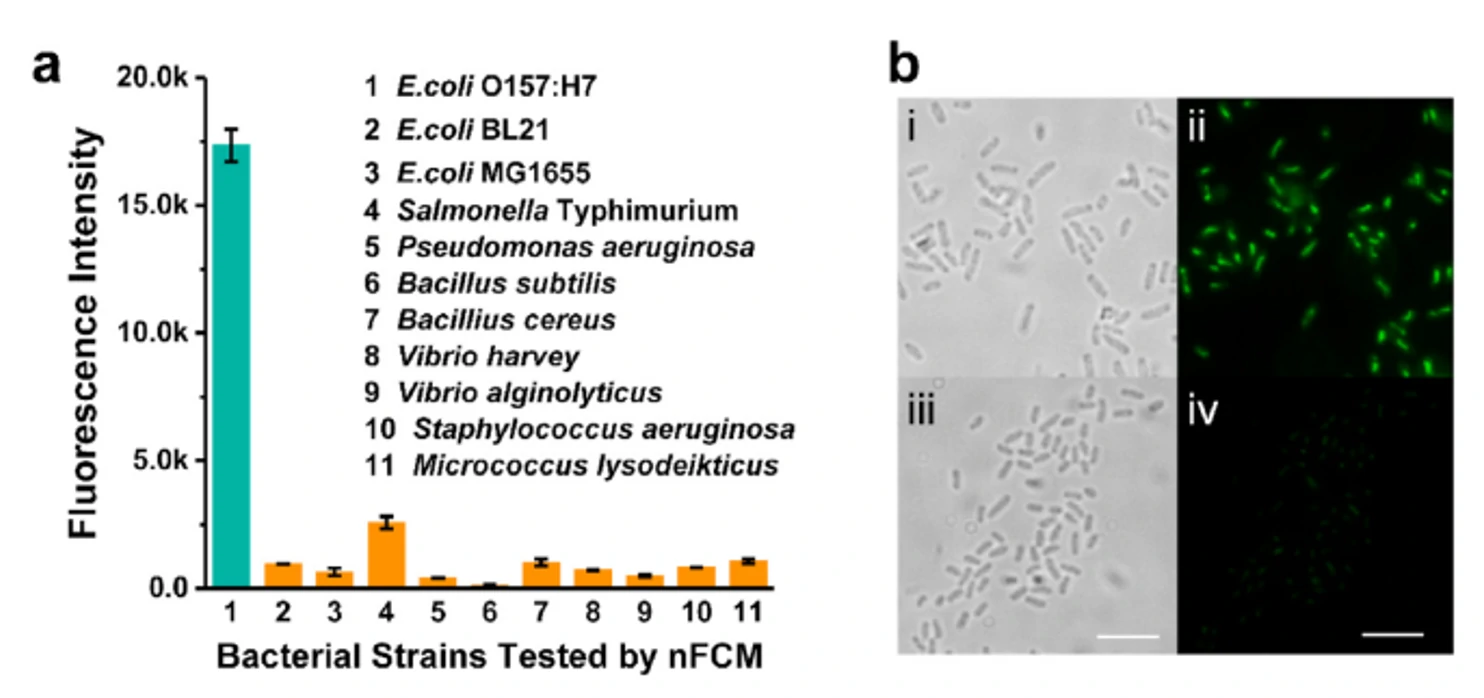
Figure 1. Binding of different strains and bacteriophages O157S-M13KE-StrB
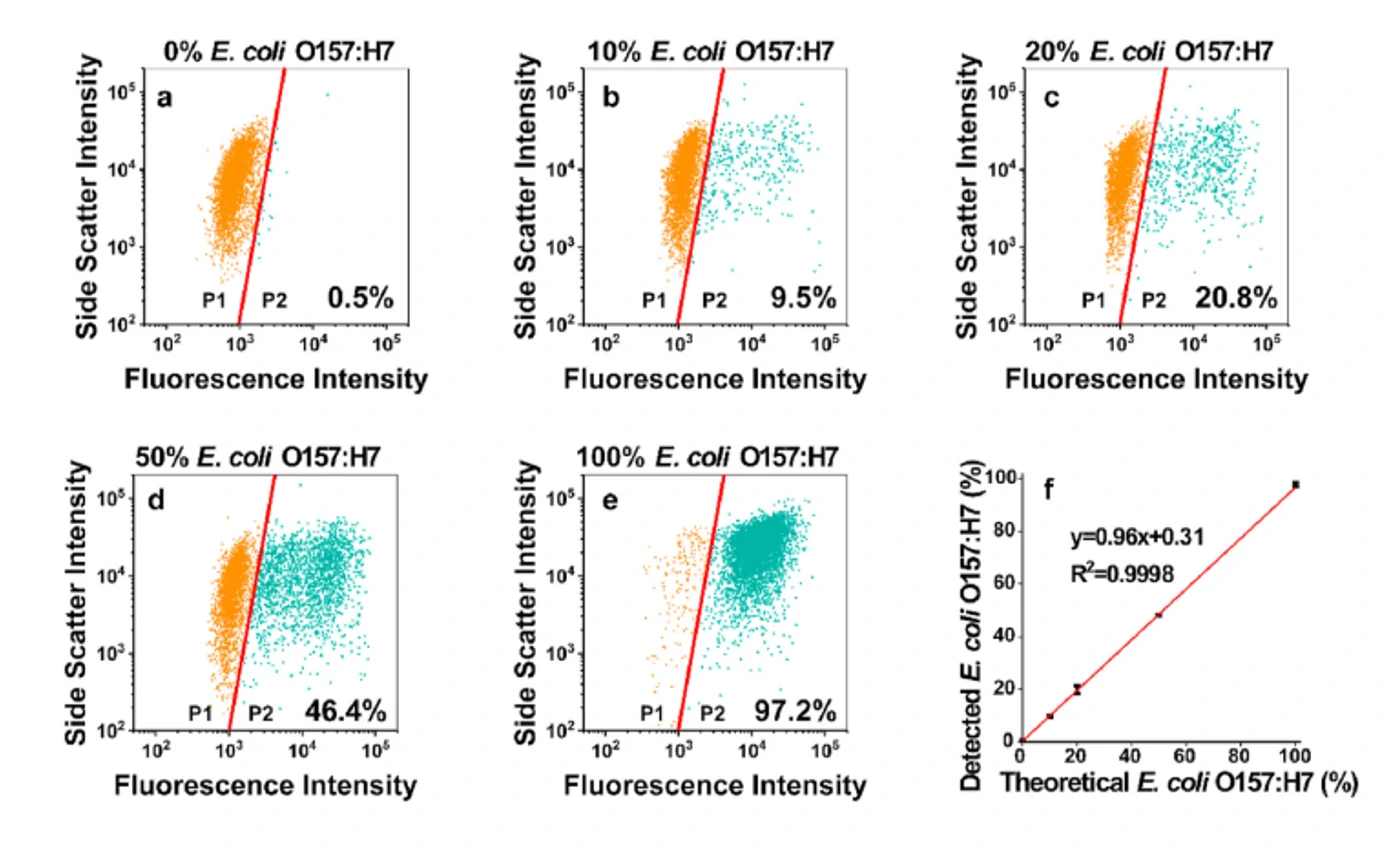
Figure 2. Mixed analysis of E. coli O157:H7 in different proportions with other non-target bacteria
The Flow NanoAnalyzer easily distinguished the strong fluorescence signal in E. coli O157:H7 and can detect weak fluorescence signals from other non-pathogenic strains.
Analytica Chimica Acta, 2021, 1166, 338596.