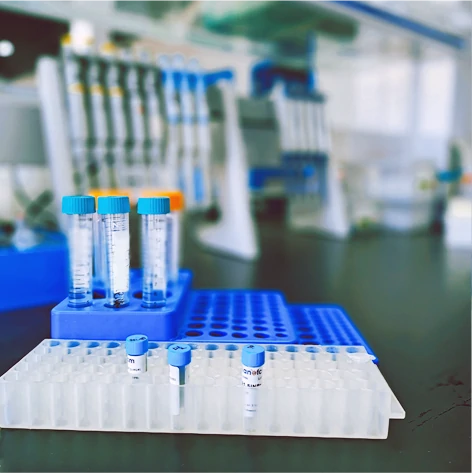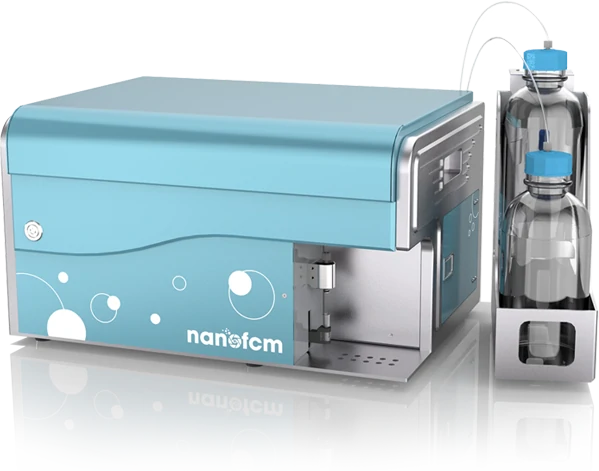
- Laser (blue, green, red)
- One side-scatter channel
- Two fluorescence channels
- up to 12,000 particles/min
- Lower detection limit 7 nm (gold nanoparticles) to 40 nm (exosomes, viruses)
- High detection limit (1 micron)
Virus Characterization for Cell and Gene Therapy with Nano Flow Cytometry :
The demand for high-purity viruses and tools for their characterization is skyrocketing with the rapid progress of cell and gene therapies. The major challenge is accurately determining the titer, structure, and contaminant levels of viral vectors throughout the development and manufacturing process. The NanoAnalzer from NanoFCM allows the multi-parameter testing of various kind of viruses, with real resolution to quantitate viral vectors in complex solutions, offering a powerful tool for cell and gene therapy industry!
Date :
Thursday, April 18, 2024
Time :
10:00 AM EST | 7:00 AM PST | 4:00 PM CEST











EXTRACELLULAR VESICLES

THE ORIGIN AND PHENOTYPING OF EXTRACELLULAR VESICLES

NANOMEDICINE

COMPREHENSIVE MEASUREMENT SUITE FOR NANOSIZED DRUG PRODUCTS

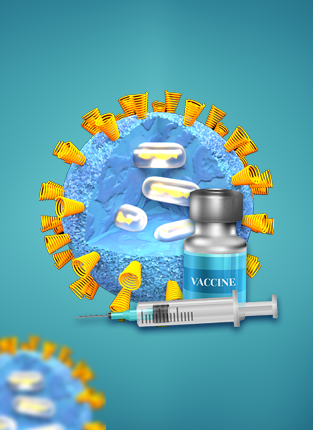
VACCINE

COMPREHENSIVE MEASUREMENT SUITE FOR NANOSIZED DRUG PRODUCTS

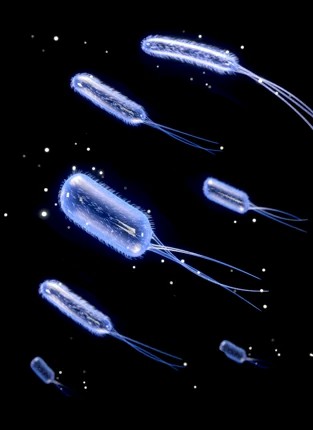
BACTERIA DETECTION

TRULY QUANTITATIVE MEASUREMENT OF TOTAL BACTERIAL TITRE

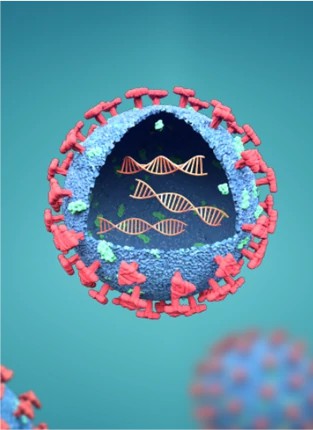
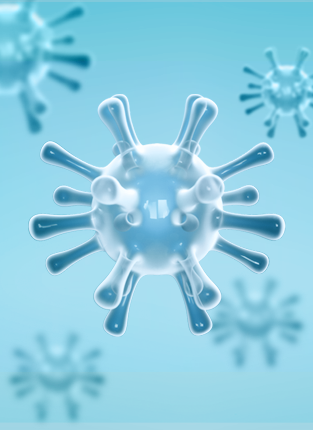
VIRUSES/VIRAL VACCINE

INTEGRATE INTO THE GENOMES OF THE HOST CELLS

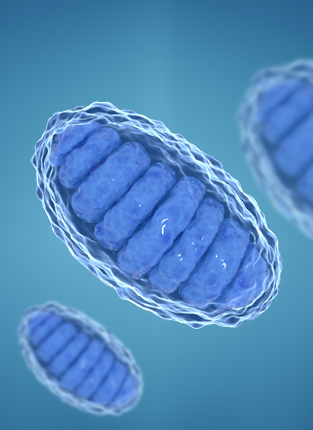
OTHERS

COMPREHENSIVE MEASUREMENT SUITE FOR NANOPARTICLES AND INDIVIDUAL MITOCHONDRIA


EXTRACELLULAR VESICLES

THE ORIGIN AND PHENOTYPING OF EXTRACELLULAR VESICLES

NANOMEDICINE

COMPREHENSIVE MEASUREMENT SUITE FOR NANOSIZED DRUG PRODUCTS

VACCINE

COMPREHENSIVE MEASUREMENT SUITE FOR NANOSIZED DRUG PRODUCTS

BACTERIA DETECTION

TRULY QUANTITATIVE MEASUREMENT OF TOTAL BACTERIAL TITRE

VIRUSES/VIRAL VACCINE

INTEGRATE INTO THE GENOMES OF THE HOST CELLS

OTHERS

COMPREHENSIVE MEASUREMENT SUITE FOR NANOPARTICLES AND INDIVIDUAL MITOCHONDRIA


High Performance


No Swarm Detection


Label Free
Flexible
Particle Concentration

Measure the absolute concentration of both total particles and subpopulations

Phenotyping

Based on single -molecule fluorescence detection, Phenotype quantify direct from marker labeling intensity.

Size Distribution


Particle Concentration
Multiparameter
Phenotyping
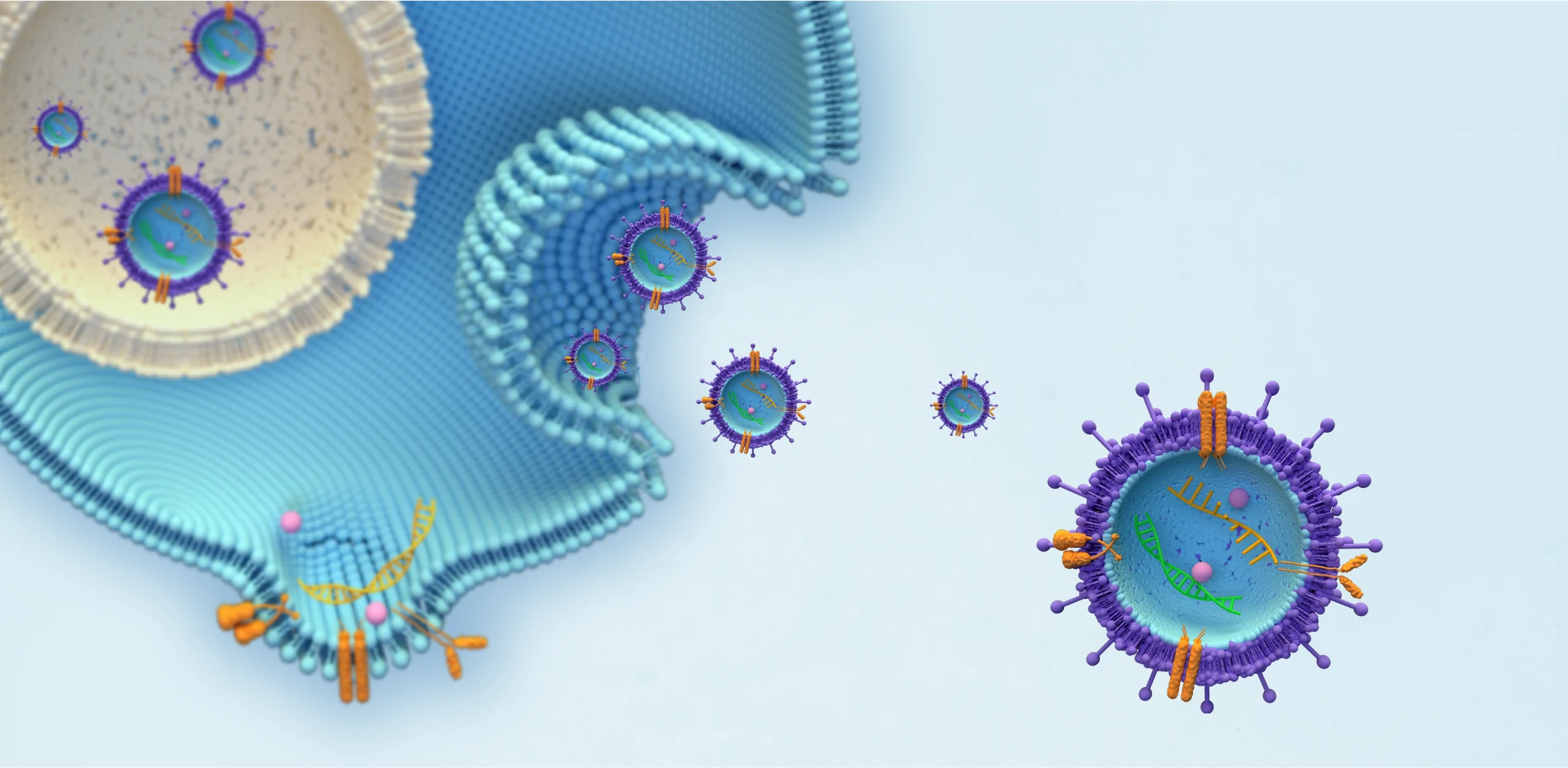
For many years scientists yearned for the possibility of performing flow cytometry to analyse small bio-nanoparticles that are too small to be measured by conventional and high sensitivity instruments. These entities, extracellular vesicles, gene therapy vectors, viruses and drug delivery particles, are promised to become the next generation of therapeutics, but they have been hard to handle and characterise due to their small size and biological or chemical heterogeneity. There is therefore a strong case for bringing flow cytometry capability to the sub-200nm scale.
NanoFCM has developed the NanoAnalyzer platform that now enables true flow-cytometry measurement at the sub-micron scale, and down to particle sizes unreachable by any other flow cytometers (10-40nm depending on the nature of the substrate). Nano-flow cytometry, the technology that underpins the NanoAnalyzer, removes bias and uncertainty stemming from the use of fluorescence signal triggering by using its highly sensitive side-scatter channel to trigger particle events. The single-particle nature of the measurement prevents uncontrolled swarming events, reinforcing data integrity. High resolution of both scatter and fluorescence channels allows the assessment of subpopulations, based on size or on bio-chemical properties.

Nano-flow cytometry’s ability to measure simultaneously a (bio)-nanoparticle population for size, size distribution and bio-chemical properties on a single instrument dramatically improves data quality and throughput compared to the traditional, multiple-techniques approach involving particle characterisation and counting (DLS, NTA, RPS), combined with chemical and biological function assessment (ELISA, Western Blot, Flow Cytometry, PCR). Quantitative measurement of the active and contaminant particles in a single preparation opens up the possibility of characterisation-based nanomedicine regulatory approval, and allows the conduct of large-scale clinical studies. From the research laboratory to the quality control department, NanoFCM delivers comprehensive bio-nano analysis.